Virtual Embryo Atlas
Developmental timeline
TS1 (fertilized egg) → P56 (adult) · click a stage to load samples

TS1
E0.5
1-cell

TS2
E1
2-cell
TS3
E2
4-16 cell
TS4
E3
morula → blastocyst

TS5
E4
late blastocyst

TS6
E4.5
attaching blastocyst
 1
1TS7
E5
egg cylinder
 1
1TS8
E6
prestreak
 1
1TS9
E6.5
streak
 2
2TS10
E7
amnion
 2
2TS11
E7.5
neural plate
 5
5TS12
E8
neural folds
 2
2TS13
E8.5
5-8 somites
 2
2TS14
E9
9-12 somites
 10
10TS15
E9.5
13-20 somites
 6
6TS16
E10
forelimb buds
 4
4TS17
E10.5
hindlimb buds
 7
7TS18
E11
handplate
 11
11TS19
E11.5
hindlimb handplate
 3
3TS20
E12
digital rays
 3
3TS21
E13.5
finger separation
 2
2TS22
E14
eyelid formation
 16
16TS23
E15
eyelids closing
 6
6TS24
E15.5
ear pinna
 5
5TS25
E16.5
long whiskers
 2
2TS26
E17.5
body hair
 1
1TS27
E18.5
pre-birth

P7
P7
postnatal day 7

P56
P56
adult (8 weeks)
E0
E2
E4
E6
E8
E10
E12
E14
E16
E18
Birth
E0E5E10E15BirthP7P56
TS19
E11.5· Hindlimb handplateGenital tubercle, vibrissae primordia. Cardiac septation underway.
8samples
2/8anatomy delineated
OPT · 8Optical Projection Tomography — a 3D scan of a chemically cleared embryo, captured by rotating it through 360° in a wide-field microscope. Isotropic voxels; raw bg = dark, tissue = bright. Stereo-seq · 2Stereo-seq (BGI) — DNB-array spatial transcriptomics with subcellular ~500 nm resolution. Captures whole-organ to whole-embryo sections; in the Spateo Cell 2024 atlas, 74 sections per embryo were aligned into a 3D cell-bin volume. sci-RNA-seq3 · 1sci-RNA-seq3 — high-throughput single-nucleus combinatorial-indexing RNA-seq (Shendure lab). The OMG mouse prenatal time-lapse profiled 11.4M nuclei across 45 developmental bins from E8 to P0 with this protocol.
Transcriptomics data · 3
Stage-matched · not registered to atlas anatomy Stereo-seq
e11_5_embryo
6,993,667 cells · 21,717 genes
Spatial · mouse · stage TS19
Qiu et al., Cell 2024 (Spateo)
Open viewer →
Stereo-seq
e11_5_heart
98,966 cells · 19,746 genes
Spatial · mouse · stage TS19
Qiu et al., Cell 2024 (Spateo)
Open viewer →
 sci-RNA-seq3
sci-RNA-seq3Mouse prenatal time-lapse · whole atlas
11,441,407 cells · 45,525 genes (full atlas)
Single-cell · pre-filtered to TS19 · spans 16 stages
Qiu et al., Nature 2024 (OMG)
Open UMAP for TS19 →
3D reference models
Histology plates · 58
Kaufman atlas, EMAPA-annotated · scroll → 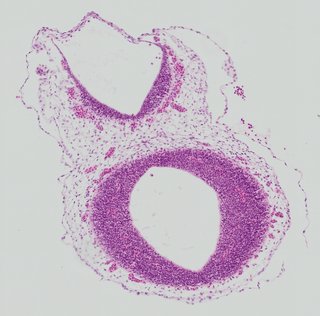 Transverse
Transverse Plate 26
TS19
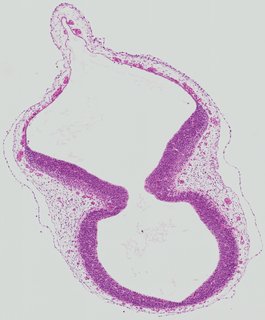 Transverse3 ann
Transverse3 ann Plate 26b
TS19
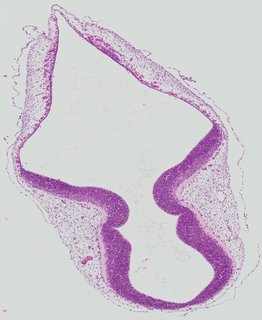 Transverse6 ann
Transverse6 ann Plate 26c
TS19
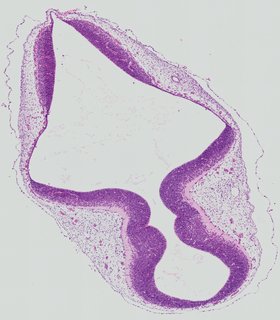 Transverse5 ann
Transverse5 ann Plate 26d
TS19
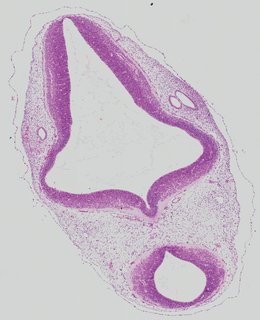 Transverse3 ann
Transverse3 ann Plate 26e
TS19
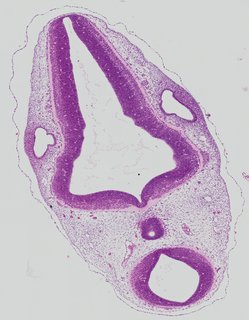 Transverse6 ann
Transverse6 ann Plate 26f
TS19
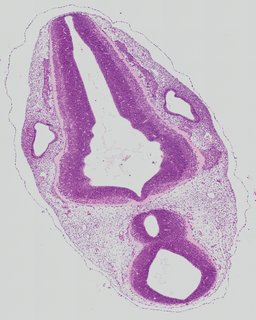 Transverse5 ann
Transverse5 ann Plate 26g
TS19
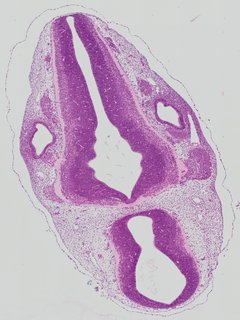 Transverse8 ann
Transverse8 ann Plate 26h
TS19
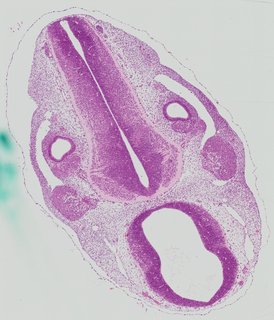 Transverse8 ann
Transverse8 ann Plate 26i
TS19
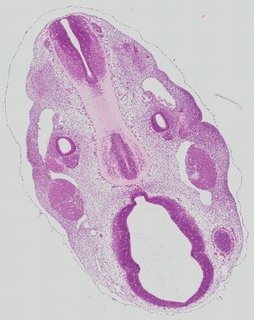 Transverse18 ann
Transverse18 ann Plate 26j
TS19
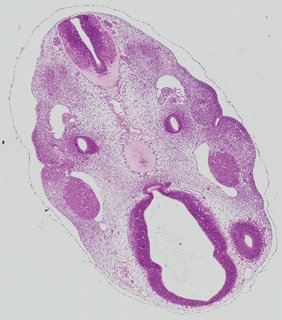 Transverse26 ann
Transverse26 ann Plate 26a
TS19
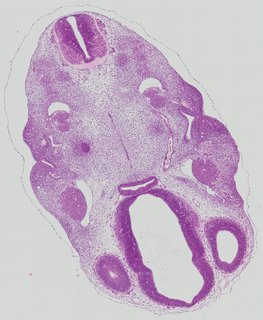 Transverse11 ann
Transverse11 ann Plate 26b
TS19
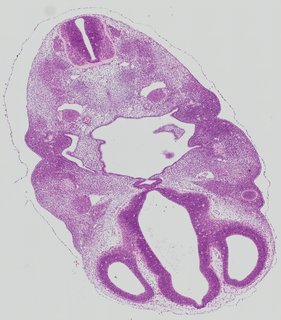 Transverse13 ann
Transverse13 ann Plate 26c
TS19
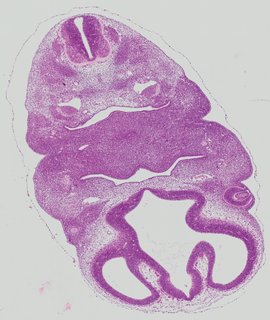 Transverse9 ann
Transverse9 ann Plate 26d
TS19
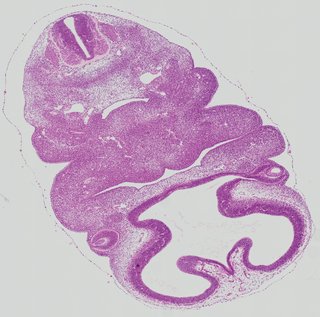 Transverse13 ann
Transverse13 ann Plate 26e
TS19
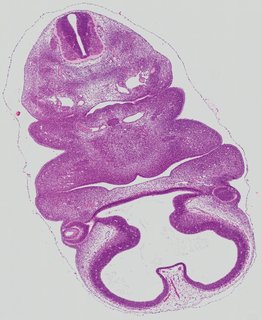 Transverse9 ann
Transverse9 ann Plate 26f
TS19
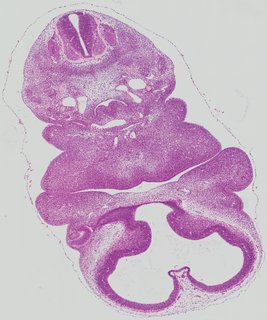 Transverse13 ann
Transverse13 ann Plate 26g
TS19
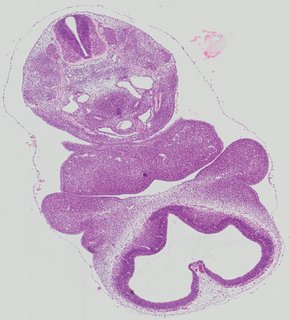 Transverse8 ann
Transverse8 ann Plate 26h
TS19
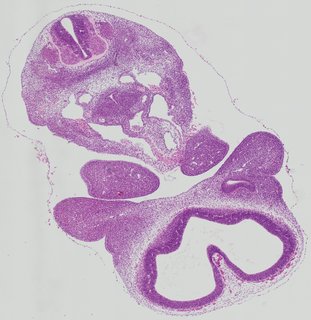 Transverse8 ann
Transverse8 ann Plate 26i
TS19
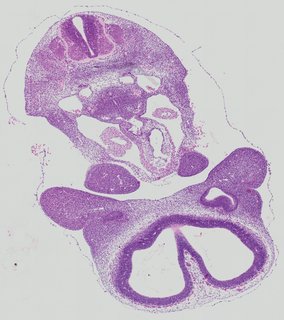 Transverse27 ann
Transverse27 ann Plate 26a
TS19
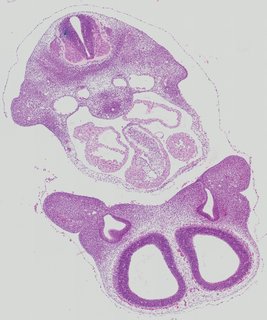 Transverse14 ann
Transverse14 ann Plate 26b
TS19
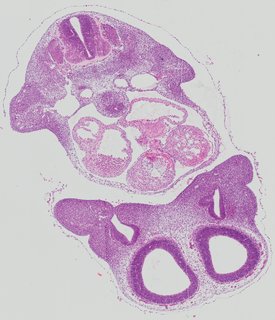 Transverse4 ann
Transverse4 ann Plate 26c
TS19
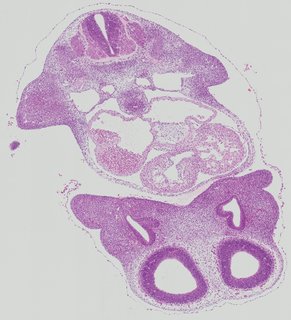 Transverse4 ann
Transverse4 ann Plate 26d
TS19
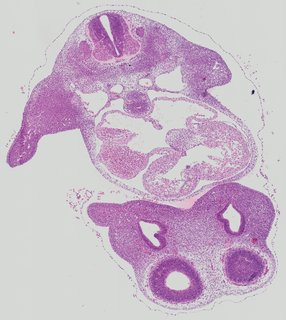 Transverse7 ann
Transverse7 ann Plate 26e
TS19
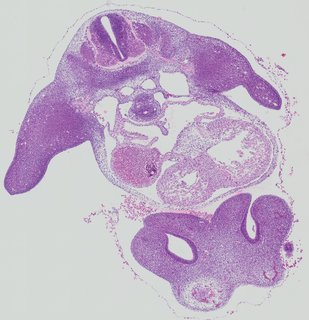 Transverse7 ann
Transverse7 ann Plate 26f
TS19
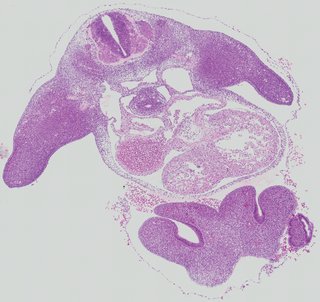 Transverse7 ann
Transverse7 ann Plate 26g
TS19
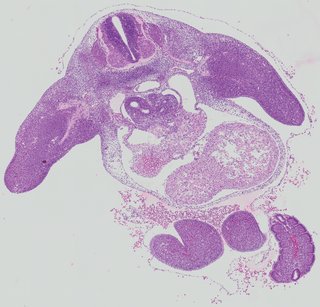 Transverse13 ann
Transverse13 ann Plate 26h
TS19
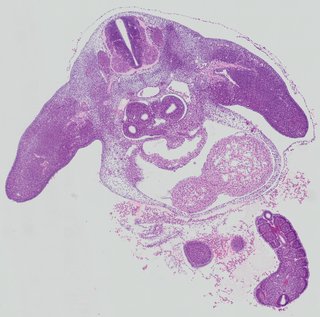 Transverse10 ann
Transverse10 ann Plate 26i
TS19
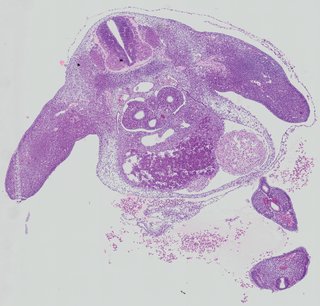 Transverse28 ann
Transverse28 ann Plate 26a
TS19
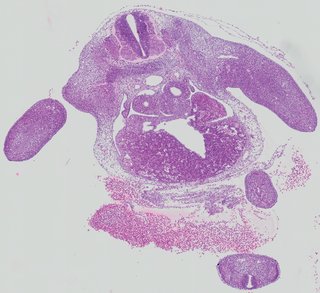 Transverse13 ann
Transverse13 ann Plate 26b
TS19
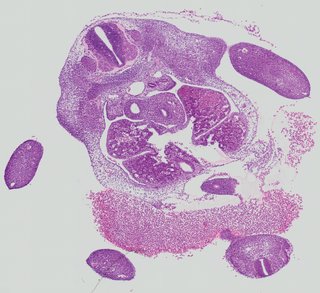 Transverse9 ann
Transverse9 ann Plate 26c
TS19
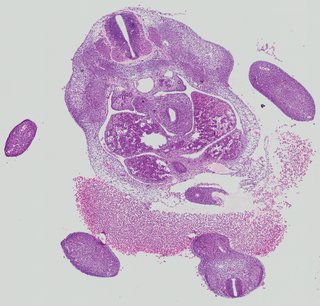 Transverse9 ann
Transverse9 ann Plate 26d
TS19
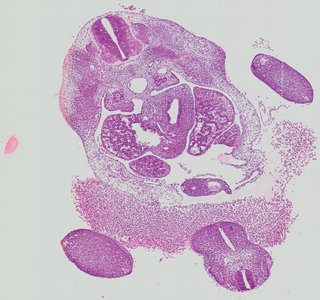 Transverse9 ann
Transverse9 ann Plate 26e
TS19
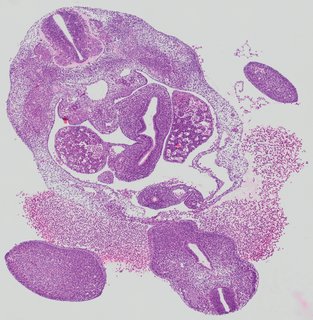 Transverse5 ann
Transverse5 ann Plate 26f
TS19
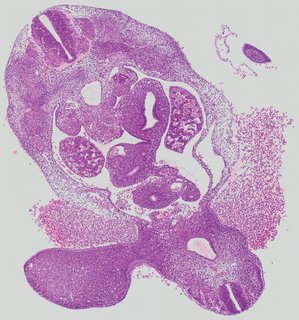 Transverse7 ann
Transverse7 ann Plate 26g
TS19
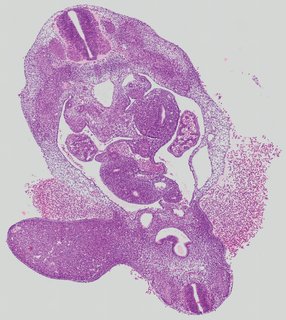 Transverse7 ann
Transverse7 ann Plate 26h
TS19
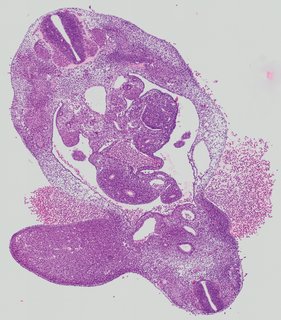 Transverse9 ann
Transverse9 ann Plate 26i
TS19
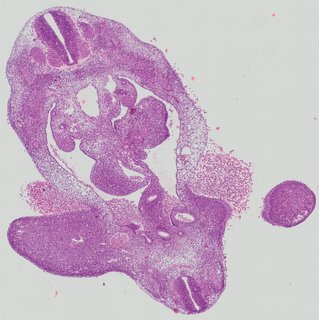 Transverse21 ann
Transverse21 ann Plate 26a
TS19
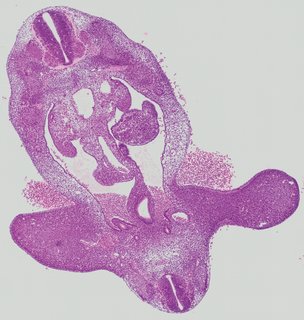 Transverse8 ann
Transverse8 ann Plate 26b
TS19
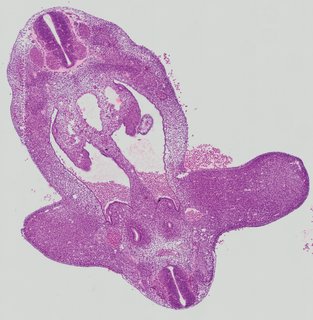 Transverse7 ann
Transverse7 ann Plate 26c
TS19
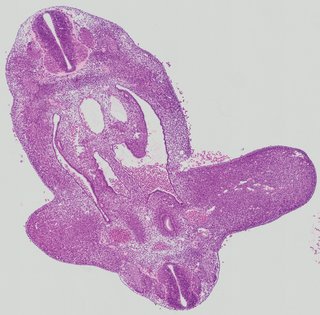 Transverse5 ann
Transverse5 ann Plate 26d
TS19
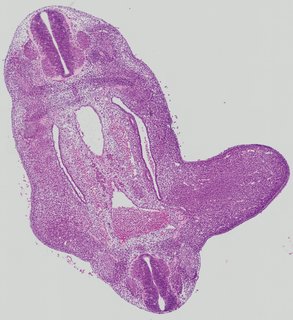 Transverse8 ann
Transverse8 ann Plate 26e
TS19
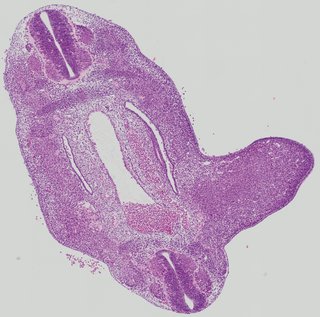 Transverse4 ann
Transverse4 ann Plate 26f
TS19
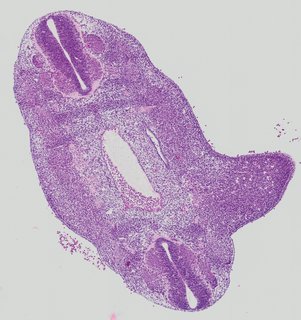 Transverse2 ann
Transverse2 ann Plate 26g
TS19
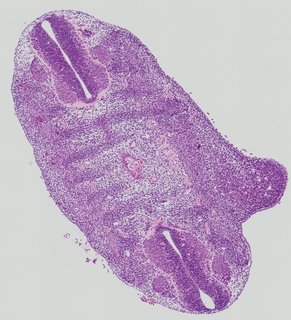 Transverse7 ann
Transverse7 ann Plate 26h
TS19
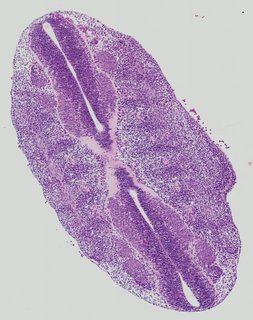 Transverse4 ann
Transverse4 ann Plate 26i
TS19
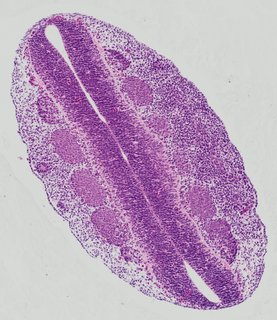 Transverse5 ann
Transverse5 ann Plate 26j
TS19
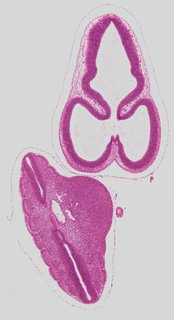 Coronal15 ann
Coronal15 ann Plate s2a
TS19
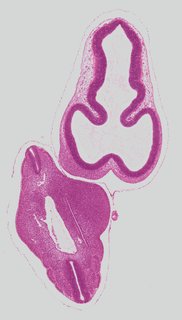 Coronal10 ann
Coronal10 ann Plate s2b
TS19
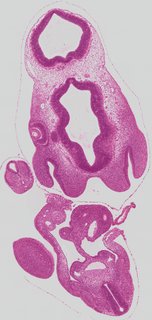 Coronal20 ann
Coronal20 ann Plate s2c
TS19
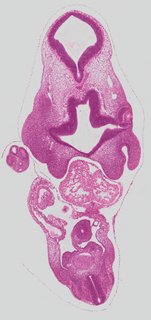 Coronal20 ann
Coronal20 ann Plate s2d
TS19
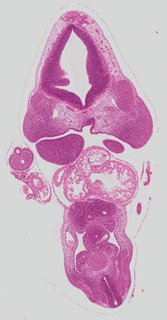 Coronal19 ann
Coronal19 ann Plate s2a
TS19
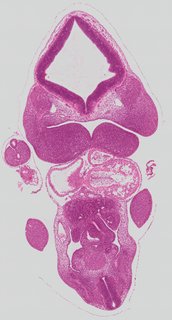 Coronal20 ann
Coronal20 ann Plate s2b
TS19
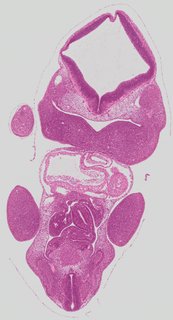 Coronal17 ann
Coronal17 ann Plate s2c
TS19
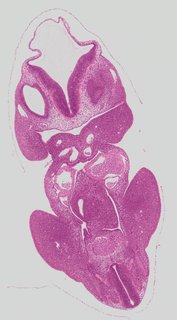 Coronal19 ann
Coronal19 ann Plate s2d
TS19
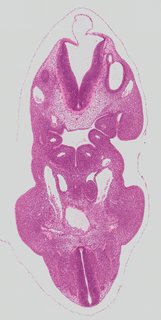 Coronal17 ann
Coronal17 ann Plate s2a
TS19
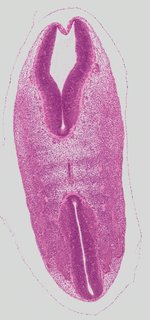 Coronal17 ann
Coronal17 ann Plate s2b
TS19
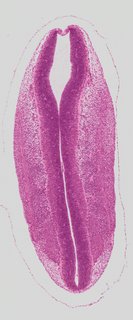 Coronal10 ann
Coronal10 ann Plate s2c
TS19
scRNA-seq · Literature · Predictions — coming soon

Previous
TS18E11
Forelimb handplate
About Theiler staging
Karl Theiler's staging system (1972) partitions prenatal development into 28 morphologically distinct steps. Virtual Embryo aligns all imagery, anatomy, and future spatial transcriptomics data to this axis.
Next
E12TS20
Digital condensations
 →
→