Virtual Embryo Atlas
Developmental timeline
TS1 (fertilized egg) → P56 (adult) · click a stage to load samples

TS1
E0.5
1-cell

TS2
E1
2-cell
TS3
E2
4-16 cell
TS4
E3
morula → blastocyst

TS5
E4
late blastocyst

TS6
E4.5
attaching blastocyst
 1
1TS7
E5
egg cylinder
 1
1TS8
E6
prestreak
 1
1TS9
E6.5
streak
 2
2TS10
E7
amnion
 2
2TS11
E7.5
neural plate
 5
5TS12
E8
neural folds
 2
2TS13
E8.5
5-8 somites
 2
2TS14
E9
9-12 somites
 10
10TS15
E9.5
13-20 somites
 6
6TS16
E10
forelimb buds
 4
4TS17
E10.5
hindlimb buds
 7
7TS18
E11
handplate
 11
11TS19
E11.5
hindlimb handplate
 3
3TS20
E12
digital rays
 3
3TS21
E13.5
finger separation
 2
2TS22
E14
eyelid formation
 16
16TS23
E15
eyelids closing
 6
6TS24
E15.5
ear pinna
 5
5TS25
E16.5
long whiskers
 2
2TS26
E17.5
body hair
 1
1TS27
E18.5
pre-birth

P7
P7
postnatal day 7

P56
P56
adult (8 weeks)
E0
E2
E4
E6
E8
E10
E12
E14
E16
E18
Birth
E0E5E10E15BirthP7P56
TS23
E15· Eyelids closingBody coat begins. Liver hematopoiesis peaks.
15samples
3/15anatomy delineated
OPT · 11Optical Projection Tomography — a 3D scan of a chemically cleared embryo, captured by rotating it through 360° in a wide-field microscope. Isotropic voxels; raw bg = dark, tissue = bright. histology · 4Histology — embryo physically embedded and microtomed into 1–7 µm sections, stained (Nissl / H&E), imaged, and reassembled into a 3D volume. High in-plane resolution, anisotropic voxels. sci-RNA-seq3 · 1sci-RNA-seq3 — high-throughput single-nucleus combinatorial-indexing RNA-seq (Shendure lab). The OMG mouse prenatal time-lapse profiled 11.4M nuclei across 45 developmental bins from E8 to P0 with this protocol.
Transcriptomics data · 1
Stage-matched · not registered to atlas anatomy 3D reference models
OPT
EMA105
· kidney
0.34×0.34×7 μm
Open viewer →
histology
EMA106
3×3×3 μm
Open viewer →
histology
EMA107
3×3×3 μm
Open viewer →
OPTmeshlabels
EMA108
· forelimb
×× μm
Open viewer →
OPTmeshlabels
EMA109
· hindlimb
×× μm
Open viewer →
histologymeshlabels
EMA147
80×80×80 μm
Open viewer →
histology
EMA80
×× μm
Open viewer →
OPT
EMA81
17.5×17.5×17.5 μm
Open viewer →
OPT
EMA82
17.5×17.5×17.5 μm
Open viewer →
OPT
EMA83
17.5×17.5×17.5 μm
Open viewer →
OPT
EMA84
17.5×17.5×17.5 μm
Open viewer →
OPT
EMA85
17.5×17.5×17.5 μm
Open viewer →
OPT
EMA86
17.5×17.5×17.5 μm
Open viewer →
OPT
EMA87
17.5×17.5×17.5 μm
Open viewer →
OPT
EMA88
17.5×17.5×17.5 μm
Open viewer →
Histology plates · 80
Kaufman atlas, EMAPA-annotated · scroll → 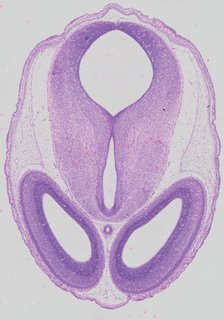 Transverse20 ann
Transverse20 ann Plate 32a
TS22 · TS23
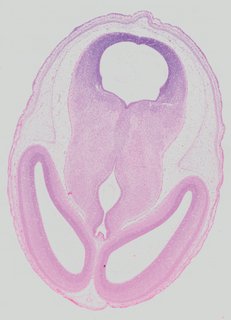 Transverse6 ann
Transverse6 ann Plate 32b
TS22 · TS23
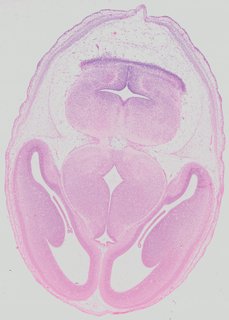 Transverse15 ann
Transverse15 ann Plate 32c
TS22 · TS23
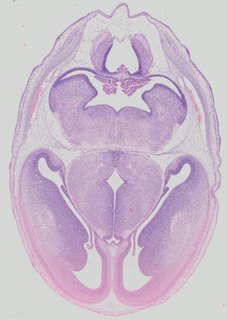 Transverse21 ann
Transverse21 ann Plate 32d
TS22 · TS23
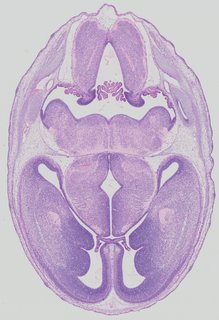 Transverse37 ann
Transverse37 ann Plate 32a
TS22 · TS23
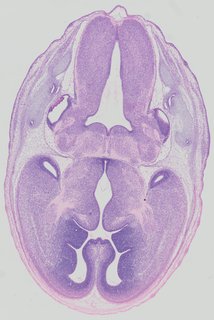 Transverse17 ann
Transverse17 ann Plate 32b
TS22 · TS23
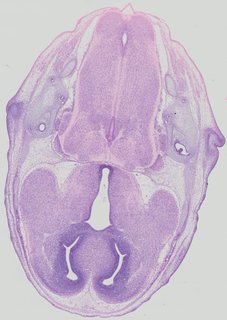 Transverse13 ann
Transverse13 ann Plate 32c
TS22 · TS23
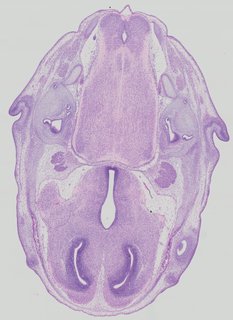 Transverse13 ann
Transverse13 ann Plate 32d
TS22 · TS23
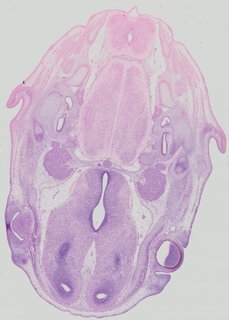 Transverse44 ann
Transverse44 ann Plate 32a
TS22 · TS23
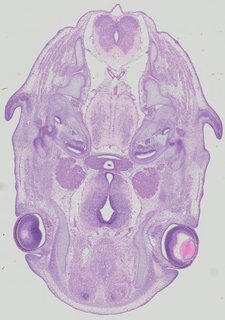 Transverse17 ann
Transverse17 ann Plate 32b
TS22 · TS23
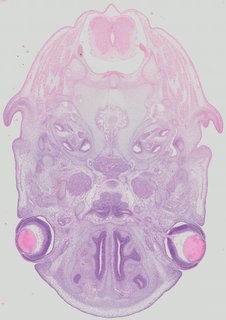 Transverse23 ann
Transverse23 ann Plate 32c
TS22 · TS23
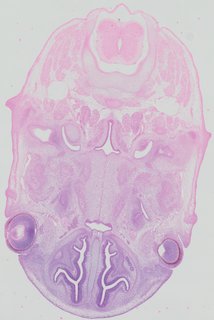 Transverse43 ann
Transverse43 ann Plate 32a
TS22 · TS23
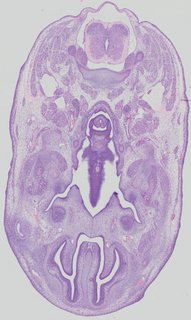 Transverse13 ann
Transverse13 ann Plate 32b
TS22 · TS23
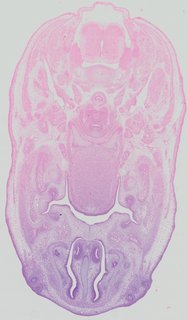 Transverse17 ann
Transverse17 ann Plate 32c
TS22 · TS23
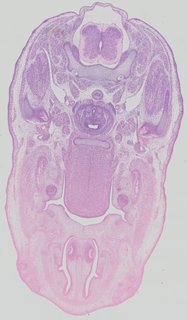 Transverse14 ann
Transverse14 ann Plate 32d
TS22 · TS23
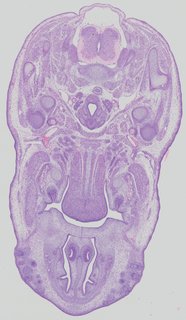 Transverse43 ann
Transverse43 ann Plate 32a
TS22 · TS23
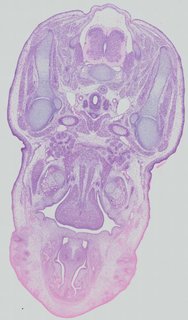 Transverse20 ann
Transverse20 ann Plate 32b
TS22 · TS23
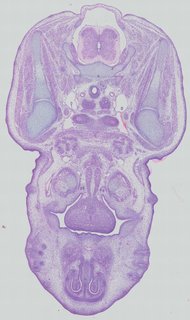 Transverse16 ann
Transverse16 ann Plate 32c
TS22 · TS23
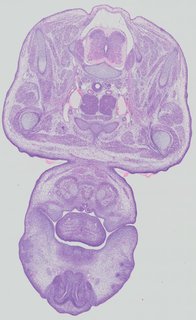 Transverse15 ann
Transverse15 ann Plate 32d
TS22 · TS23
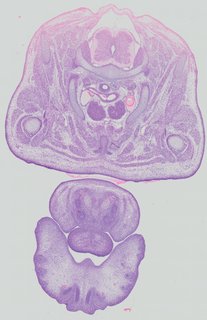 Transverse41 ann
Transverse41 ann Plate 32a
TS22 · TS23
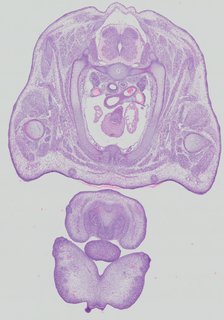 Transverse18 ann
Transverse18 ann Plate 32b
TS22 · TS23
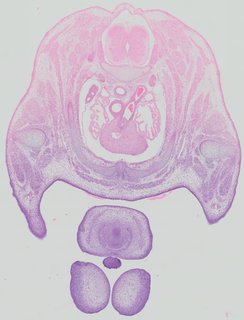 Transverse13 ann
Transverse13 ann Plate 32c
TS22 · TS23
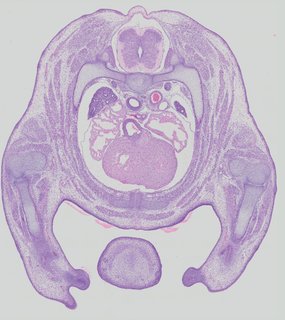 Transverse24 ann
Transverse24 ann Plate 32d
TS22 · TS23
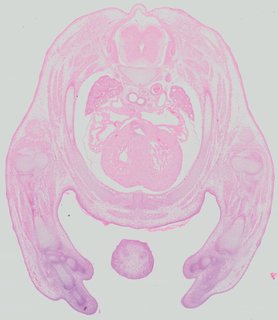 Transverse42 ann
Transverse42 ann Plate 32a
TS22 · TS23
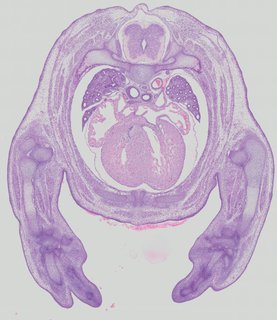 Transverse23 ann
Transverse23 ann Plate 32b
TS22 · TS23
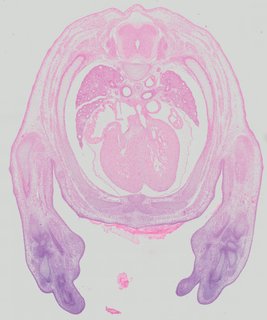 Transverse12 ann
Transverse12 ann Plate 32c
TS22 · TS23
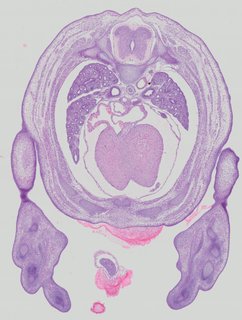 Transverse15 ann
Transverse15 ann Plate 32d
TS22 · TS23
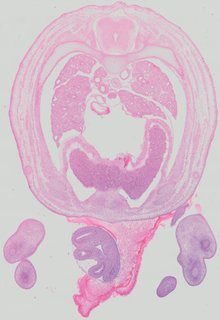 Transverse36 ann
Transverse36 ann Plate 32a
TS22 · TS23
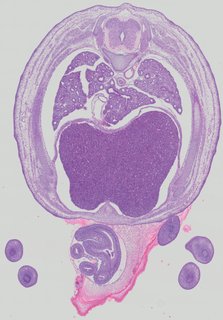 Transverse14 ann
Transverse14 ann Plate 32b
TS22 · TS23
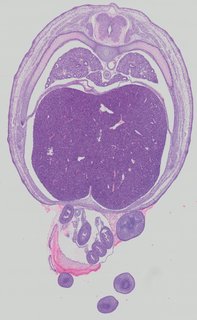 Transverse9 ann
Transverse9 ann Plate 32c
TS22 · TS23
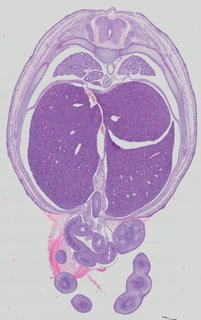 Transverse13 ann
Transverse13 ann Plate 32d
TS22 · TS23
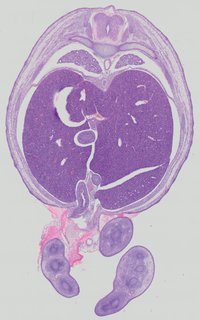 Transverse39 ann
Transverse39 ann Plate 32a
TS22 · TS23
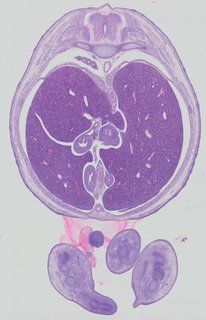 Transverse16 ann
Transverse16 ann Plate 32b
TS22 · TS23
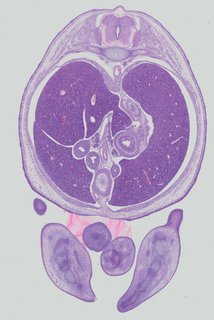 Transverse13 ann
Transverse13 ann Plate 32c
TS22 · TS23
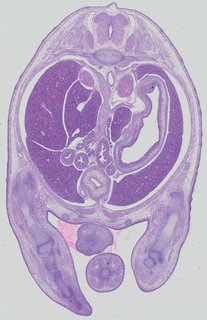 Transverse20 ann
Transverse20 ann Plate 32d
TS22 · TS23
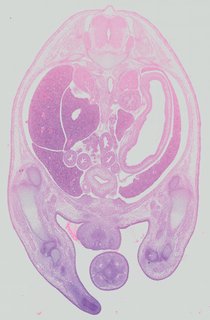 Transverse47 ann
Transverse47 ann Plate 32a
TS22 · TS23
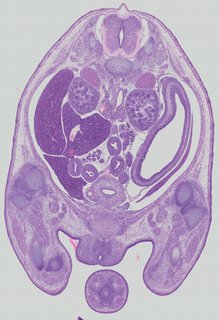 Transverse9 ann
Transverse9 ann Plate 32b
TS22 · TS23
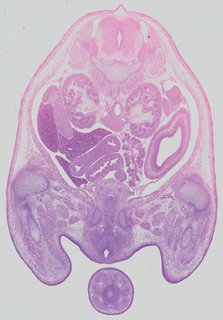 Transverse16 ann
Transverse16 ann Plate 32c
TS22 · TS23
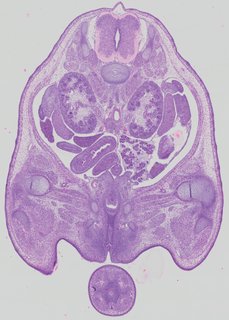 Transverse16 ann
Transverse16 ann Plate 32d
TS22 · TS23
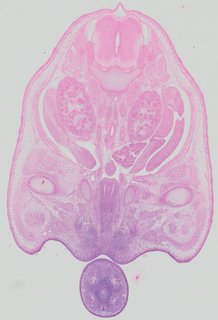 Transverse43 ann
Transverse43 ann Plate 32a
TS22 · TS23
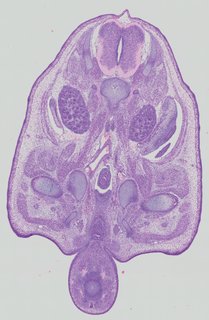 Transverse15 ann
Transverse15 ann Plate 32b
TS22 · TS23
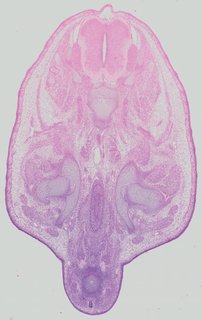 Transverse12 ann
Transverse12 ann Plate 32c
TS22 · TS23
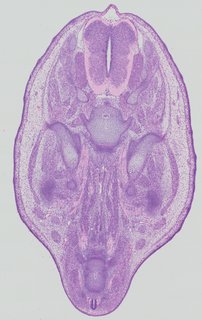 Transverse5 ann
Transverse5 ann Plate 32d
TS22 · TS23
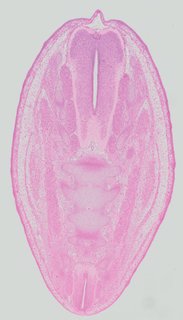 Transverse3 ann
Transverse3 ann Plate 32e
TS22 · TS23
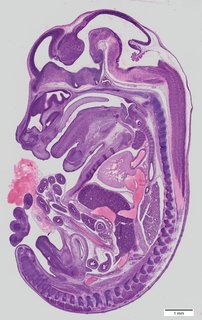 Sagittal76 ann
Sagittal76 ann Plate 33a
TS22 · TS23
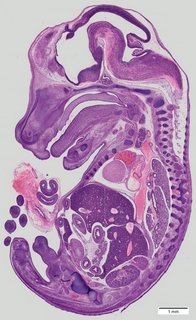 Sagittal41 ann
Sagittal41 ann Plate 33b
TS22 · TS23
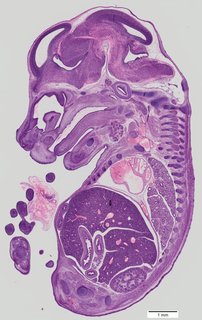 Sagittal76 ann
Sagittal76 ann Plate 33a
TS22 · TS23
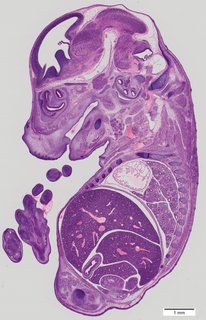 Sagittal38 ann
Sagittal38 ann Plate 33b
TS22 · TS23
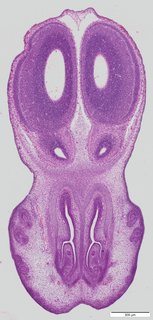 Coronal13 ann
Coronal13 ann Plate 34a
TS22 · TS23
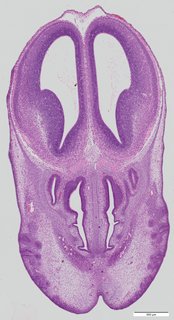 Coronal10 ann
Coronal10 ann Plate 34b
TS22 · TS23
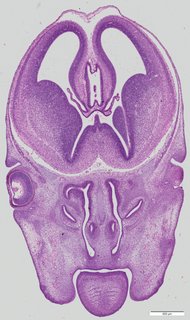 Coronal15 ann
Coronal15 ann Plate 34c
TS22 · TS23
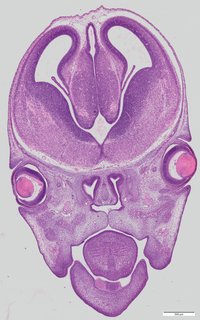 Coronal17 ann
Coronal17 ann Plate 34d
TS22 · TS23
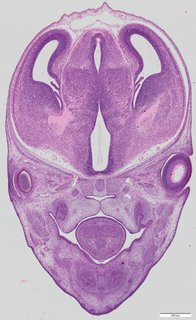 Coronal14 ann
Coronal14 ann Plate 34e
TS22 · TS23
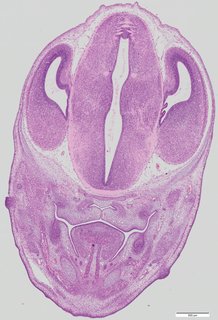 Coronal16 ann
Coronal16 ann Plate 34f
TS22 · TS23
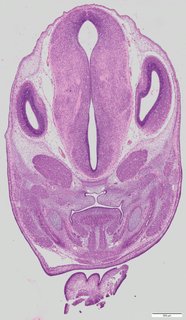 Coronal8 ann
Coronal8 ann Plate 34g
TS22 · TS23
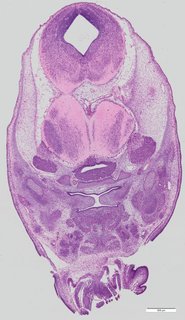 Coronal15 ann
Coronal15 ann Plate 34h
TS22 · TS23
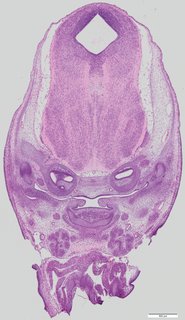 Coronal18 ann
Coronal18 ann Plate 34a
TS22 · TS23
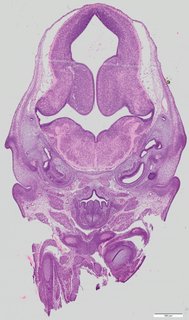 Coronal19 ann
Coronal19 ann Plate 34b
TS22 · TS23
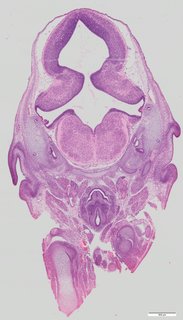 Coronal10 ann
Coronal10 ann Plate 34c
TS22 · TS23
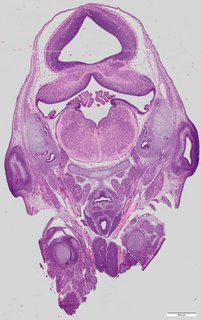 Coronal16 ann
Coronal16 ann Plate 34d
TS22 · TS23
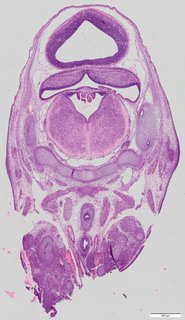 Coronal7 ann
Coronal7 ann Plate 34e
TS22 · TS23
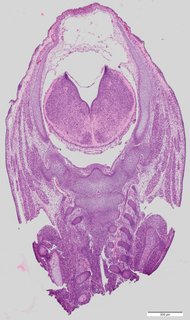 Coronal10 ann
Coronal10 ann Plate 34f
TS22 · TS23
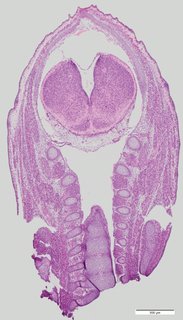 Coronal7 ann
Coronal7 ann Plate 34g
TS22 · TS23
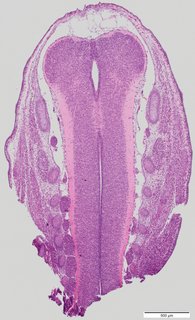 Coronal5 ann
Coronal5 ann Plate 34h
TS22 · TS23
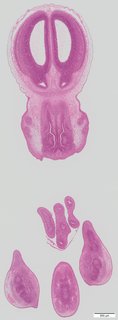 Coronal14 ann
Coronal14 ann Plate s5a
TS23
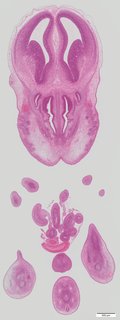 Coronal16 ann
Coronal16 ann Plate s5b
TS23
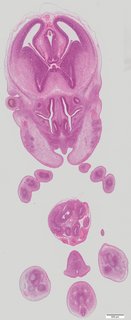 Coronal21 ann
Coronal21 ann Plate s5c
TS23
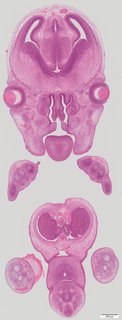 Coronal25 ann
Coronal25 ann Plate s5d
TS23
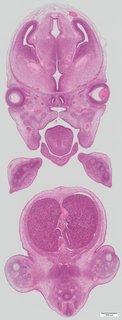 Coronal26 ann
Coronal26 ann Plate s5a
TS23
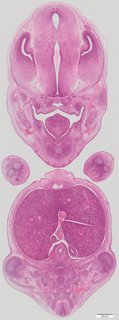 Coronal20 ann
Coronal20 ann Plate s5b
TS23
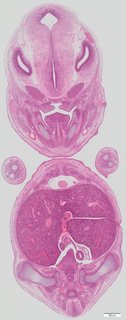 Coronal16 ann
Coronal16 ann Plate s5c
TS23
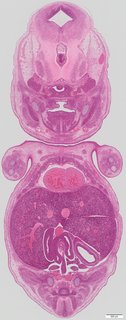 Coronal13 ann
Coronal13 ann Plate s5d
TS23
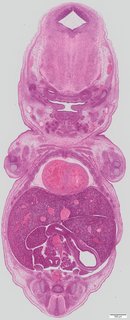 Coronal20 ann
Coronal20 ann Plate s5a
TS23
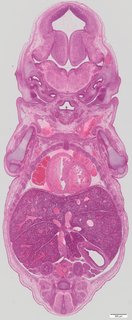 Coronal21 ann
Coronal21 ann Plate s5b
TS23
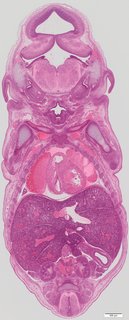 Coronal21 ann
Coronal21 ann Plate s5c
TS23
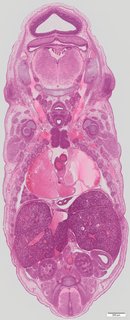 Coronal33 ann
Coronal33 ann Plate s5d
TS23
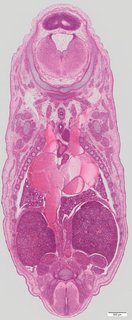 Coronal22 ann
Coronal22 ann Plate s5a
TS23
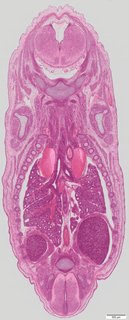 Coronal19 ann
Coronal19 ann Plate s5b
TS23
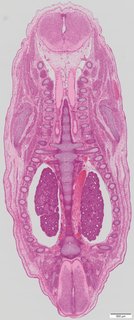 Coronal18 ann
Coronal18 ann Plate s5c
TS23
 Coronal13 ann
Coronal13 ann Plate s5d
TS23
scRNA-seq · Literature · Predictions — coming soon

Previous
TS22E14
Eyelids begin
About Theiler staging
Karl Theiler's staging system (1972) partitions prenatal development into 28 morphologically distinct steps. Virtual Embryo aligns all imagery, anatomy, and future spatial transcriptomics data to this axis.
Next
E15.5TS24
Ear pinna
 →
→